Morphological pseudotime ordering and fate mapping reveals diversification of cerebellar inhibitory interneurons
Abstract
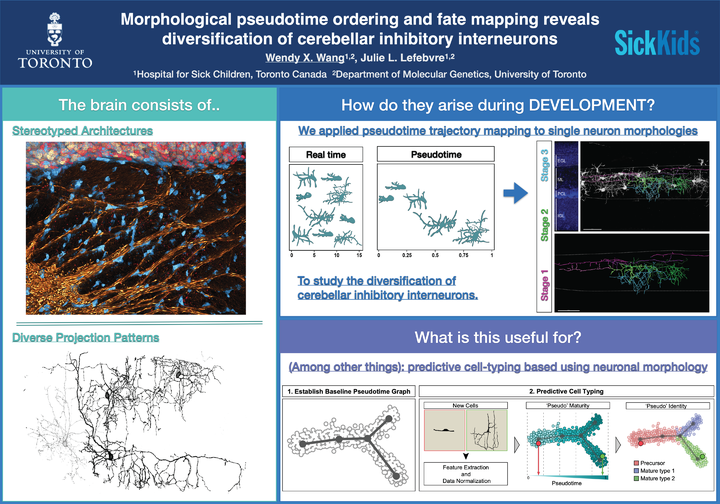
Understanding how diverse neurons are assembled into circuits requires a framework for describing cell types and their developmental trajectories. Here, we combined genetic fate mapping and pseudo-temporal profiling to resolve the diversification of cerebellar inhibitory interneurons based on morphology. The molecular layer interneurons (MLIs) derive from a common progenitor but comprise a diverse population of dendritic-, somatic-, and axon initial segment-targeting interneurons. MLIs are classically divided into two types. However, their morphological heterogeneity suggests an alternate model of one continuously varying population. Through clustering and trajectory inference of 811 MLI reconstructions at maturity and during development, we show that MLIs divide into two discrete classes but also present significant within-class heterogeneity. Pseudotime trajectory mapping uncovered the emergence of distinct phenotypes during migration and axonogenesis, well before neurons reach their final positions. Our study illustrates the utility of quantitative single-cell methods to morphology for defining the diversification of neuronal subtypes.